In the previous post we looked at some of the initial results of the YFULL analysis of the Big Y test from two of the volunteers from Genetic Family 1 (GF1). In this post we will take a closer look at the SNPs revealed by the additional YFULL analysis and then compare and contrast them with the original results from FTDNA.
This is quite a long post but stick with it!
Terminal SNPs
The two volunteers from GF1 were given new ID numbers at YFULL and you can see them in the haplotree diagram below - they are the last two numbers.
- The first volunteer (FTDNA kit number 164729) is YF04104 (results available in Sep 2015)
- The second volunteer (H1223) is YF04316 (results Oct 2015)
 |
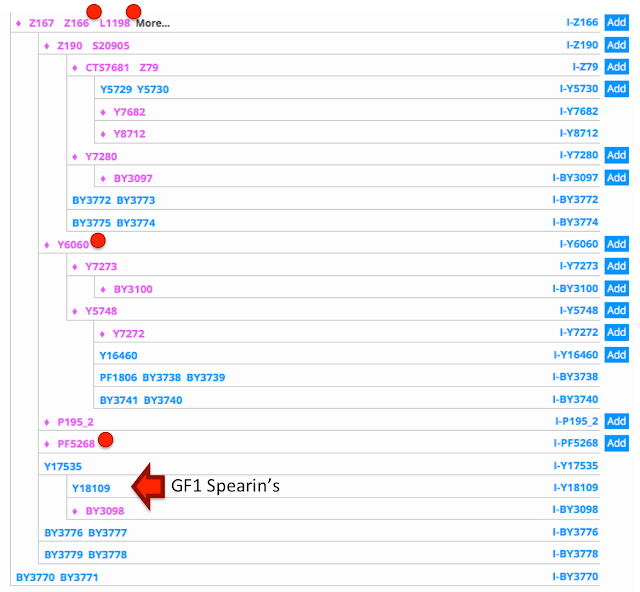
| The terminal SNP Block for GF1 on the YFULL Haplotree |
Our two brave volunteers have been placed on the
YFULL Haplotree as a sub-branch below SNP Y17535. Both our volunteers have the terminal SNP Y18109 or rather they have a whole "block" of terminal SNPs, namely:
- Y18109
- Y18110
- Y18111
- Y18112
- Y18113
- Y18114
- Y18115
- Y18116
- Y18117
- Y18118
Shared SNPs
When two or more people share several terminal SNPs in common, this terminal SNP "block" is usually named after the first SNP in the block, which in our case is Y18109. These terminal SNPs are shared between our two volunteers and no other people in the world (currently).
If we move up the tree to the next nearest branching point, we find this is marked by the SNP Y17535 (which represents a SNP block of 5 SNPs). There are 2 sub-branches below Y17535 - our own sub-branch (Y18109) with our own 2 volunteers, and another sub-branch (Y17535*) with a single individual. Thus 3 individuals (currently) share the SNP Y17535.
 |
Shared SNPs on the L1198 branch of the YFULL haplotree
(click to enlarge) |
And if we move further upstream to the next nearest branching point, this is marked by the SNP L1198 (representing another SNP block of about 7 SNPs, although some are "equivalent SNPs" i.e. the same SNP with several different names because it was discovered by several different people around about the same time - these are the SNPs separated by a forward slash). There are 4 sub-branches below L1198 - the Y17535 branch just discussed above (with 3 individuals), but also an L1198* sub-branch (3 people), a Y6060 sub-branch (with 3 subsequent sub-branches, the last of which also has 3 sub-branches; 9 people in total), and an S20905 sub-branch (aka Z190, but not shown in the diagram for some strange reason; with 4 levels of sub-branching) containing 11 people altogether (currently).
So, in total, 26 people share the SNP block L1198, 3 people share the SNP block Y17535, and 2 people (our volunteers) share the SNP block Y18109.
Currently.
But as more people test, either from GF1 or our close genetic neighbours (if they exist), each SNP block should be gradually split up. In other words, we can expect our particular sub-branch of the tree to be joined by adjacent sub-branches sprouting nearby, some of which will "steal" SNPs from our current terminal SNP block. We can also expect further sub-branches to sprout below our current terminal SNP / SNP block. For example, if (say) 10 people from GF1 were to test, the current 10-SNP block might dwindle to (say) a 2-SNP block, with (say) 4 sub-branches below it - one sub-branch containing a single terminal SNP, another containing a 3-SNP block, and two containing a 2-SNP block.
The Take Home Message is: our current terminal SNP block will dwindle and will be split up as more people do the Big Y test (or similar tests).
Unique (Personal) SNPs
In addition to the 10 SNPs that the two volunteers share in common with each other (i.e. the Y18109 SNP Block), they each possess SNPs that the other does not have. In other words, they have their own unique, personal or "private" SNPs that no one else in the world has (currently). No doubt if we were to test other members of GF1 for these "private" SNPs we would find that some of these unique SNPs would no longer be unique anymore - they would be shared with other members of the Spearin group - thus dwindling the number of unique SNPs possessed by any given individual.
YFULL reports that Member YF04316 (H1223) has 16 "Novel SNPs" with 3 SNPs characterised as Best Quality, 1 as Acceptable Quality, and 11 as Ambiguous Quality. These are illustrated in the diagram below. In contrast, member YF04104 (164729) has 51 Novel SNPs but none are of best quality, 1 is of acceptable quality, 49 are of ambiguous quality, and 1 is of low quality (these are not shown here because they take up too much space). But what do they mean by quality?
The quality of a SNP is a reflection of how confident the company is about declaring it to be a true positive SNP and not a false positive finding. There are various reasons for why the test might throw up a false positive result and we don't need to go into the details here, but it is simply important to remember that some results may be false positives and it is best to focus on the SNPs that the company is most confident about (i.e. the best quality SNPs).
Unique SNPs (currently) possessed by member YF04316
If more people from GF1 tested, we would probably find that some of the 16 Best Quality Novel SNPs of Member YF04316 (H1223) would turn up in the results of some of the new people, and would no longer be "private" or unique - they would be shared by other members in the group. And this might even result in one or several more branches being formed.
So, in a similar way to how the shared SNPs in the current GF1 terminal SNP block will split up as our genetic neighbours get tested, these unique SNPs to H1223 will also gradually disappear as more people test. So, for example, if
everyone from GF1 were to do the Big Y test, a lot of H1223's unique SNPs would turn out to be shared by other members of GF1 (and thus they would not be unique any more). This could be useful when building a
Mutation History Tree (discussed in a subsequent blog post) but we could also probably achieve this with the existing STR data instead, so there is no burning need for more people in GF1 to do the Big Y test.
Comparison between FTDNA Analysis & YFULL Analysis
We have looked at the YFULL reanalysis of the Big Y data. Now we are going to compare it to the Big Y data analysis originally performed by FTDNA to see if (and where) there are similarities and differences.
The FTDNA results report that our two volunteers match on 24,165
known SNPs and differ on 2 known SNPs, namely YSC0000155 and PF3643. In fact it is member 164729 who appears to be lacking these SNPs - H1223 appears to have them both. This is very surprising given that we expect our two volunteers to be related by a common ancestor some time in the 1600's and so there should be a very close relationship between them with no major differences in the SNPs they share. So for them to differ by two SNPs is quite a surprise.
Furthermore, these two SNPs in question are nowhere to be found on either the FTDNA Haplotree or the
ISOGG Haplotree. I found YSC0000155 on
YBrowse and it was discovered in a Haplogroup J-L147 person but there is no further information available on this SNP. Similarly, PF3643 was discovered in 2011 and possibly belongs in Haplogroup I. The
I-M223 Yahoo Discussion Group notes that PF3643 turns up in some but not all I-M223 people and that "
some people's Big Y test did not record a result for PF3643. However, there is enough data to show that Z79+ people must have had a back mutation from derived C back to ancestral A." So it is difficult to judge whether these SNPs are relevant to our own particular Spearin sub-branch of the human evolutionary tree. I suspect that these particular SNPs may be quite far upstream from where we currently sit and are of no particular relevance to the conversation that follows. But I could be wrong.
Furthermore, the nature of NGS tests (Next Generation Sequencing tests) like the Big Y means that this particular test simply failed to detect these two SNPs this time around and they are in fact present after all. If we were to repeat the same test in the same individual they might pop up in the second test.
A big thank you to John Cleary who pointed out that you can check SNP information on YFULL if you know the SNP name. Just go to Check SNPs, enter the name of the SNP in question and click on the magnifying glass icon when it comes up.
I was able to check the YFULL website for FTDNA's mystery missing Known SNPs (YSC0000155 and PF3643) but obtained no additional useful information. I still do not know where these are placed in the haplotree. Perhaps they have not been allocated a position as yet.
 |
Enter a SNP name to get SNP details
(click to enlarge) |
But the above discussion relates to "known" SNPs. Let's take a look at the "unknown SNPs - the "Novel" SNPs.
Shared SNPs
According to the FTDNA analysis, our two volunteers have 201 "Shared Novel Variants" but when you click on the number 201, the pop-up box not only has Shared Novel Variants but also the SNPs unique to each of the two individuals. So this should not really be under the heading "
Shared Novel Variants" as it also includes "unique" variants that are not shared with anyone. A relatively minor criticism, but potentially confusing.
 |
| FTDNA's Big Y results page for H1223 - 201 "Shared Novel Variants" with 164729 |
There are 3 tabs in the Shared Novel Variants pop-up box - one tab has 156 "Shared" SNPs, 45 "unique" to H1223, and 13 "unique" to 164729 ... and that adds up to a total of 214 ... so where does the 201 come from?? 156 + 45 is 201 ... so did they forget the other 13 SNPs? Other numbers for nearby neighbours (155, & 190) also
do not add up correctly. This is not potentially confusing - it
is confusing.
 |
| Pop-up box with 3 tabs showing Shared SNPs & unique SNPs |
Apart from the confusion over the term "Shared" and the actual number of SNPs detected, there are several further sources of confusion.
Firstly, the definition of the term "
Novel" in the phrase "Shared Novel Variant". Novel is supposed to refer to SNPs that have never been discovered before. But ... before when? The definition of Novel varies between companies so what is novel to FTDNA may not be considered novel to YFULL. And vice versa. Furthermore, presumably anything "novel" has a time-limit, after which it becomes classified as "known" ... but no one knows when this time-limit expires. And this may also differ among companies ... one man's "cutting edge" may be another's "yesterday's news". There is no standardisation. So caution is necessary when interpreting these results and comparing them between companies. There will be differences in how companies report the same data.
Here's another source of confusion. FTDNA reports 156 Shared SNPs whereas YFULL does not give this actual number - it places the two individuals together on the YFULL tree sharing 10 SNPs in their shared Terminal SNP Block (Y18109), 5 SNPs shared at the branching point above that (Y17535 branch), and possibly 7 SNPs on the branching point above that (L1198 branch). So,
where on the tree are these 156 shared SNPs that FTDNA says the two volunteers share? Do they go right back up the tree, back to "genetic Adam"?
And this is also where we encounter our next problem - FTDNA do not report SNP names,
only SNP positions. This makes it difficult to identify SNPs and compare results between companies - some people use SNP names for identification, other people use SNP positions. In order to find out the SNP names (and thereafter ascertain where on the tree they sit), we have to enter every SNP position on
YBowse to see if there is a corresponding name (or several corresponding names). That's 156 SNP positions!! What a pulaver!
Below is a screenshot of
ISOGG's YBrowse utility. By entering the position in the search box, you can find if there are any particular SNPs at that particular position on the Y chromosome. You have to enter the position in the format shown. The example below is for position 7,321,330 and there are (apparently) 4 different SNP names at this particular position. This initially suggests that they are all
equivalent SNPs (i.e. same SNP, different names) but further examination of the Details for each of the 4 SNPs reveals that there is a
contradictory direction of mutation - was it from C to A (SNPs 1,3,4), or from A to C (SNP 2)? Which came first? The chicken (C) or the Egg (A)? [Note:
allele-anc refers to the ancestral value (i.e. the original or reference value) and
allele-der refers to the derived or mutated value.]
 |
| Browse reveals there are 4 SNPs at position 7,321,330 |
 |
Details of the 4 SNPs with contradictory directions of mutation
(click to enlarge) |
A further point of confusion is the fact that this particular
SNP is found in several Haplogroups, namely R, O & Q, whereas we know the Spearin's are in Haplogroup I. So ... what does this mean? This does not look like a SNP that is uniquely shared by just our two volunteers. It appears to be a SNP that is shared not just by our two volunteers but by a host of other people??... including people in other haplogroups? In which case, there is really not much point in me trying to identify all 156 "shared SNPs" that FTDNA says our two volunteers have in common.
I stopped after five!
What about the 10 SNPs shared between our two volunteers (the so-called Y19108 block) on the YFULL tree? Are these included in FTDNA's list of 156 shared SNPs? And what about the shared SNPs further upstream at branching points (Y17535, L1198, etc)? Are these also in the FTDNA list of shared SNPs?
Well, it was possible to use
YBrowse to identify the positions for each of the SNPs on the YFULL tree. And then compare these positions to the FTDNA list to see if they appeared there. Here's what was found:
- all 7 SNPs in the L1198 block are missing from FTDNA's Shared Novel Variants list ... but this could be because they are relatively well-established "upstream" SNPs and therefore do not meet the criteria for "Novel"
- 3 of the 5 SNPs in the Y17535 block are present in FTDNA's list but 2 are missing (see diagram below) ... however, one of them (Y17491) turns up in FTDNA's list of unique SNPs for H1223 (YF04316)! It seems this particular SNP was recognised as a unique SNP by FTDNA but as a shared SNP by YFULL. So who is "right"?
- 6 of the 10 SNPs in the Y18109 block are present in FTDNA's list but 4 are missing (Y18109, -10, -16, & -18) ... and again, 2 of them turn up in FTDNA's list of unique SNPs for H1223. These SNPs are identified as unique by FTDNA but shared by YFULL. So who do we believe?
The fact that the Y18109 SNP is missing from FTDNA's Shared SNP list is highly confusing because FTDNA have assigned the terminal SNP for both our volunteers as Y18109. How can they do this if it does not turn up as a shared SNP between the two volunteers??? However it does appear in the list of SNPs tested for each of our volunteers on their Haplotree & SNPs page on the FTDNA website. And when I download the SNPs from each volunteer into a csv file, there it is, Y18109, in both files, and derived from the Big Y test! Why then does it not turn up in the Shared Novel Variants list? Perhaps it is classified as a "known" SNP? And that's why it turns up in the downloaded csv file but the others do not? But that still does not explain the absence of the other 3 missing SNPs from our terminal Y18109 SNP block.
It's a conundrum. A quandary. A mystery.
 |
| A portion of my spreadsheet with the 156 Shared Novel Variants reported by FTDNA |
 |
| Only 3 of the 5 SNPs in the Y17535 SNP Block appear on FTDNA's Shared Novel Variants list |
So FTDNA do not identify all the shared SNPs identified by YFULL. Possibly because the two companies have different thresholds / criteria for declaring a SNP to be present.
But it points to a
major lack of consistency between the YFULL analysis and the FTDNA analysis. And this naturally will raise concerns in people's minds about the accuracy of the data. Who got it right? Maybe both companies did. Maybe the differences are all down to the different criteria employed by each company for declaring a SNP. Or maybe not. Which analysis do you believe? Which is more reliable?
And what about the rest of the 156 Shared SNPs? Only 9 SNPs relate to the 3 branches of the YFULL tree discussed above - where do the other 147 fit in? Are they further upstream?
It would be much more helpful if FTDNA simply reported the SNPs shared uniquely by Person A and Person B and no one else.
So, thus far, the analysis of FTDNA's 156 Shared SNPs has not been very helpful at all. Maybe we'll have better luck with the unique SNPs?
Unique SNPs
FTDNA reports that H1223 (YF04316) has 45 unique SNPs (i.e. not shared with
164729 / YF04104
) and similarly 164729 (YF04104) has 13 unique SNPs (i.e. not shared with H1223 / YF04316). This differs considerably from the 16 and 51 unique SNPs reported by YFULL above.
 |
| Unique SNPs reported by each company |
But once again, the different companies have different criteria for declaring a SNP and this effects the results. If we take a closer look at the reporting criteria, FTDNA describe their "confidence" in the SNP as high, medium or unknown. In contrast, YFULL describes the "quality" of the SNP as best, acceptable, ambiguous, & low. Neither set of criteria are right or wrong - merely different approaches.
And when we compare the two sets of unique SNPs, there is only agreement between FTDNA and YFULL with regard to 2 unique SNPs for member H1223 (YF04316) and 1 unique SNP for member 164729 (YF04104). These are illustrated in the diagrams at the end of this post.
- Note that for H1223, none of YFULL's "Ambiguous quality" SNPs are reported by FTDNA. And similarly all but 2 of FTDNA's "high confidence" SNPs are reported by YFULL. There are 3 "Best Quality" SNPs from YFULL (green and yellow highlight) but only 2 of these (yellow highlight) are declared by FTDNA.
- For 164729 (YF04104), only 1 unique SNP is declared by both companies (yellow highlight). This is deemed to be of "high confidence" by FTDNA and "acceptable quality" by YFULL.
Therefore, in terms of consistency or agreement between the two companies, the vast majority of unique SNPs declared by one company are not declared by the other. In terms of percentages this works out as: 2/45 (4.4%) and 1/13 (7.7%) agreement for FTDNA; and 2/16 (12.5%) and 1/51 (2%) for YFULL. This gives an average consistency score of a mere 6.7%. Or to put it another way, the companies will disagree 93.3% of the time.
Conclusions
So, even though we have a huge amount of information from both analyses, there are major differences between the two companies and what they put in their reports. The amount of inconsistency is quite astounding and highlights the need for caution in interpreting these reports.
To resolve these inconsistencies in reporting, we have to delve deeper into the data itself. And that means exploring the vcf files, bed files, and BAM files that contain the fine details of our DNA results (not accessible to Project Administrators without the express permission of the project members concerned). This is not a job for the faint-hearted and involves many hours of review and analysis. It is not a task that most Surname Project Administrators would embrace, and personally, I leave this type of analysis to the experts - the Haplogroup Project Administrators. This highlights the need for a close collaboration with people like Wayne Roberts and Aaron Salles Torres who are administrators of the
I-M223 project. They have an overview of much more data than any Surname Project Administrator, and can potentially see patterns that would be easily missed by someone looking at a mere subset of the data.
Despite all the above caveats, we have actually learnt quite a lot from SNP testing. Both interpretations of the SNP data (by FTDNA and YFULL) place us in more or less the same position on their respective haplotrees. They both assign the same terminal SNP (Y18109). And there is some (minor) agreement on what are likely to be unique SNPs for each individual.
This entire exercise has been very useful in highlighting the fact that there is no standardisation currently in the way that the data from the Big Y test is analysed and interpreted. The same applies to other NGS tests, such as those offered by FGC (Full Genomes Corporation). And this is no surprise. We have to bear in mind that we are on the crest of the wave of scientific discovery here. We are the first explorers in a brave new world. As a community, it will take time for us to take in what we are seeing, analyse it, make sense of it, and arrive at a consensus regarding the best way to interpret and present the data. As Humphrey Bogart said to Claude Rains, this is simply the start of a wonderful relationship.
In the next post we will be looking at a topic that is (perhaps) a little bit more straightforward: TMRCA estimates - the Time to the Most Recent Common Ancestor.
Maurice Gleeson
April 2016
 |
| Unique SNPs for member H1223 - only 2 SNPs were jointly declared by both companies |
 |
| Unique SNPs for member 164729 - only 1 SNP was jointly declared by both companies |
Update 4 August 2016
I received this helpful comment from the I-M223 Yahoo Discussion Group:
Regarding the numbers reported in the Shared Novel Variants pop-up boxes I can offer the following. The first tab in a Shared Novel Variants pop-up box is shown in the attached figure. It states that there are 157 shared entries. Notice the position 14263127 is ancestral, i.e. G-G, and should not be in the list. There is one other like that so in reality there are 155 shared entries. In this case the mystery number is 200 leaving 45 that I cannot account for.
I reconciled the above against novel variants listed in the data exported as CSV files. Those same two bogus entries are present. The other kit in the comparison has 18 such bogus entries. After elimination of the bogus entries one kit has 28 novel variants not shared and the other has 26. These agree with the numbers reported as not shared in the other tabs of the pop-up box.
So from this it appears that there is a bug in the FTDNA system but this does not account for the discrepancies previously noted.